Correct option is A
Explanation:
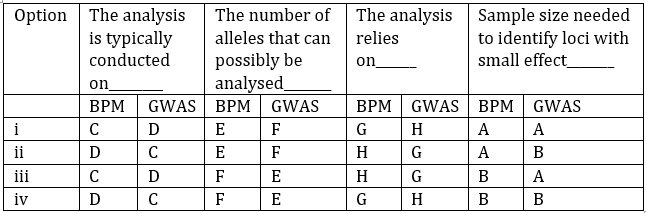
BPM is typically conducted on populations derived from controlled crosses (C) — these are experimental populations made by crossing two parents.
GWAS is typically done on random mating natural populations (D), such as landraces or natural accessions.
BPM analyzes limited two alleles per locus (E), as crosses involve only two parents.
GWAS analyzes multiple alleles per locus (F) because natural populations harbor diverse alleles.
BPM relies on linkage-based mapping (G), tracking recombination events in controlled crosses.
GWAS relies on linkage disequilibrium-based mapping (H), exploiting historical recombination in natural populations.
Both methods require a large sample size (A) to identify loci with small effects, since detecting minor QTL effects needs statistical power.
Information Booster:
Bi-parental mapping (BPM) provides high power to detect QTLs segregating between two parents but is limited in allelic diversity.
GWAS captures natural allelic diversity but requires dense markers and large sample sizes due to population structure and allele frequency.
The distinction helps breeders and geneticists choose appropriate mapping strategies.