Correct option is B
Option 2 refers to the concept of using a third marker that is located between the two genes. A third marker can help refine the recombination frequency. If the third marker lies between y and cv, it would cause the recombination frequency to increase due to additional crossovers, but it will never decrease. This is because recombination frequency can only be affected in one direction: increasing due to additional crossovers being observed. Recombination frequencies cannot decrease once the initial crossover events are accounted for.
Information Booster:
- A third marker can be used in gene mapping to help refine the distance between two genes.
- A genetic distance between genes indicates the probability of crossover events occurring between them.
- Recombination frequencies above 50% indicate independent assortment, while frequencies below 50% indicate linkage.
- The genetic distance measured in centimorgans (cM) represents the frequency of recombination events between genes.
- The addition of a third marker can reveal the position of the genes relative to each other and improve the accuracy of the genetic distance calculation.
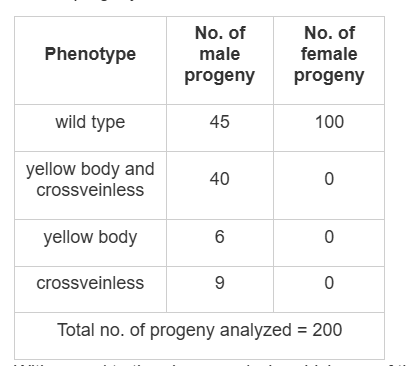
- In the case of Drosophila melanogaster, X-linked genes often show recessive inheritance patterns, as seen in the data where male progeny exhibit the traits.
- Gene mapping in Drosophila is commonly used to analyze the linkage and distance between genes on the X chromosome.
Additional Knowledge:
- Option 1 mentions a genetic distance of 7.5 cM, which is based on the initial calculation from the data provided. However, the presence of a third marker would affect the observed recombination frequency in the progeny, potentially increasing the genetic distance due to more crossovers being identified.
- Option 3 discusses progeny size, but increasing the progeny size would help detect more crossovers, it does not change the basic principle of the genetic distance, which would be increased due to the addition of a third marker.
- Option 4 refers to the reciprocal parental cross, which would suggest no recombination events, but we observe recombination here, indicating that crossover is happening in the F2 progeny.