Correct option is D
EXPLANATION-
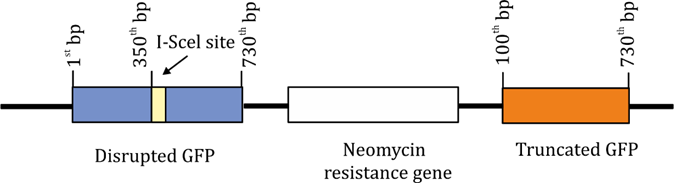
A. Gene Conversion (GC) → GFP-positive & Neomycin-resistant- Correct
GC uses a homologous template to accurately repair DSB. The right GFP (despite the 5′ deletion) can act as a template for homologous repair. If recombination spans the entire GFP region, it restores full GFP function. The neomycin resistance gene remains, so the cells would be neomycin-resistant.
B. GC → GFP-positive but Neomycin-sensitive- Incorrect
This suggests neomycin gene gets lost during repair. But in GC, no large deletions or loss of intervening sequences occur. The neomycin resistance gene is retained.
C. SSA (Single-Strand Annealing) → GFP-positive & Neomycin-resistant - Incorrect
SSA aligns homologous sequences flanking the break, deleting the region between them. This would delete the neomycin resistance gene between the GFP repeats.
So, GFP function is restored, but neomycin resistance is lost.
D. NHEJ (Non-Homologous End Joining) → GFP-negative & Neomycin-resistant- Correct
NHEJ ligates broken ends without using homology. Likely results in imprecise repair → GFP not restored.
The neomycin resistance gene remains untouched.
So, the correct option is (d) - A and D only