Correct option is D
Explanation-
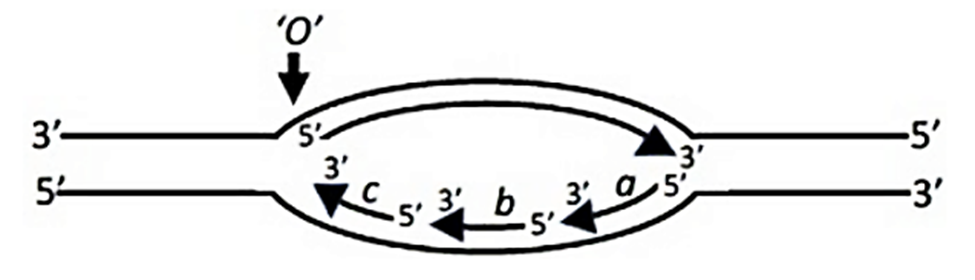
Option a: The rate of forward movement of DnaB helicase along the template DNA increases ~10-fold when DnaB and DNA Pol III interact, thus ensuring that the helicase does not move ahead rapidly without the polymerase.
Correct.
This is a known coordination mechanism in the replisome — the helicase (DnaB) and DNA polymerase III interact to synchronize their movement. When DnaB is not coupled with DNA Pol III, its activity is slower; the interaction accelerates helicase movement appropriately.
Option b: The transient interaction of the primase with the helicase allows activation of primase activity by ~1000-fold, promoting RNA primer synthesis.
Correct.
The interaction between primase and helicase is essential for primer synthesis, and this interaction indeed activates primase activity substantially (often cited as ~1000-fold). This allows timely RNA primer production.
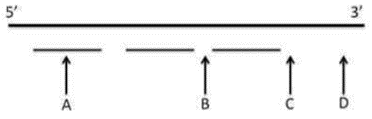
Option c: The length of the Okazaki fragments is typically restricted to 1000-2000 nucleotides.
Correct.
In prokaryotes, Okazaki fragments typically range from 1000 to 2000 nucleotides. This is a well-established fact.
Option d: The E. coli oriC carries repeats of two sequence motifs: repeats of a 9-mer that collectively form the site at which the origin first becomes single-stranded, and repeats of a 13-mer to which the DnaA initiator protein binds.
Incorrect.
This statement mixes up the function of the 9-mer and 13-mer motifs. The 9-mer sequences are binding sites for DnaA, the initiator protein. The 13-mer sequences are AT-rich regions that become single-stranded (melted) first during initiation.
So, the roles of 9-mer and 13-mer are reversed in the statement.