Correct option is C
Mismatch repair (MMR) is a crucial cellular process to correct DNA mismatches such as C:C mismatches. This process involves multiple proteins that work sequentially to detect, excise, and repair the mismatched region.
- The Msh2-Msh6 complex plays a primary role in recognizing the mismatch and recruiting downstream repair proteins.
- Polδ (DNA polymerase delta) is involved in DNA synthesis during the repair process.
- ExoI (Exonuclease I) and Rad27 (a flap endonuclease, FEN1) are involved in the excision of mismatched DNA.
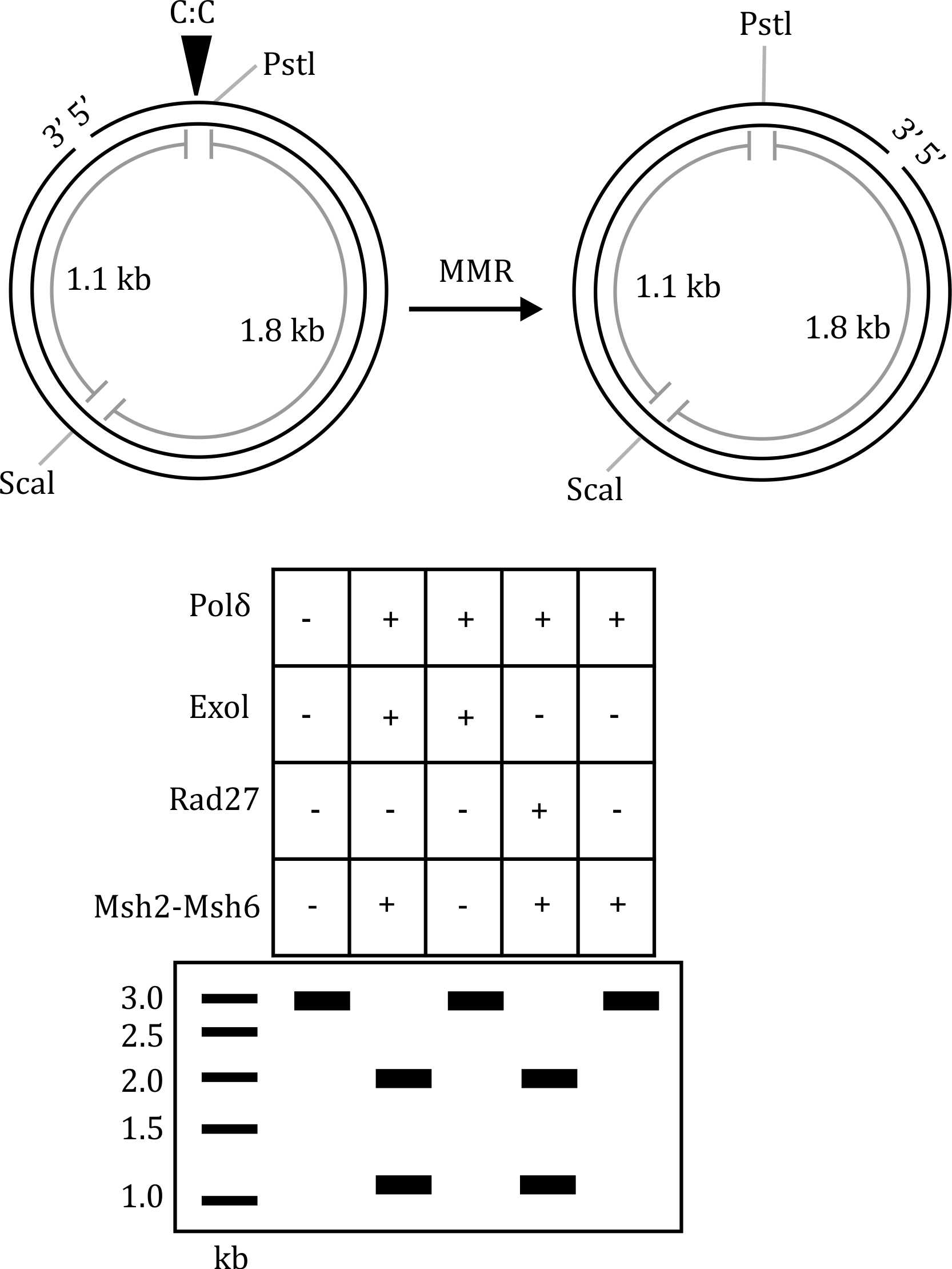
From the gel electrophoresis results:
- The presence of Msh2-Msh6, Polδ, ExoI, and Rad27 leads to successful repair, as indicated by the restoration of the PstI site and the expected digestion pattern.
- However, when ExoI and Rad27 are absent, the repair is incomplete, suggesting that Polδ and Msh2-Msh6 alone are not sufficient for C:C mismatch repair.
- This means that statement (3) is incorrect because Polδ and Msh2-Msh6 alone cannot complete the repair process without ExoI or Rad27.
Information Booster
- Mismatch Repair (MMR) Pathway: A critical pathway that maintains genome stability by correcting base mismatches arising during DNA replication.
- Msh2-Msh6 Complex: Recognizes mismatches and initiates repair by recruiting exonucleases and polymerases.
- Polδ (DNA Polymerase Delta): Synthesizes new DNA to replace the excised mismatched segment.
- ExoI and Rad27 Functions: ExoI is a 5’ to 3’ exonuclease that removes mismatched DNA, while Rad27 (FEN1) processes Okazaki fragments and assists in DNA repair.
- Requirement of Exonuclease Activity: The gel results confirm that at least one exonuclease (ExoI or Rad27) is required for successful repair.
- Functional Redundancy in DNA Repair: ExoI and Rad27 can function redundantly, meaning if one is absent, the other can still facilitate repair.
- Reconstitution Experiments: These experiments help to dissect the roles of individual proteins in DNA repair pathways.