Correct option is B
Explanation:
Given Information:
DNA Fragment A (plus strand):
5’-AGAGAGAGAGAGAGAG...GAGAGAGAGAGAGAGAGAGAG-3’DNA Fragment B (plus strand):
5’-AGCTTGCATGCCTGCAGGTCGACT---TAGACGATGA-3’
Primers:
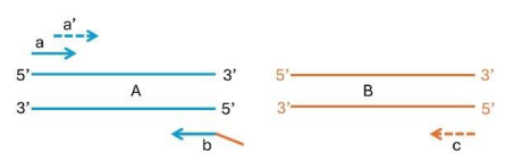
Primer “a”: Used to amplify DNA “A”.
Primer “b”: This primer is used in conjunction with primer “a” and is designed to bind to a complementary sequence in DNA “A”.
Primer “c”: This is likely used for amplifying DNA “B” in a subsequent PCR setup.
Identify the Overlapping Regions:
Since the fragments are to be stitched together, primer "b" needs to be complementary to a sequence at the end of fragment A.The overlap region would ideally have a sequence that connects the end of fragment A to the beginning of fragment B.
Determine Primer Sequence:
The sequence of primer "b" must be designed to bind to the last part of DNA "A" that is adjacent to the junction where it will connect to DNA "B".Given the requirement for proper annealing, the primer should bind at the site where DNA "B" starts, maintaining compatibility for PCR extension.
Looking at the terminal sequences of fragment A and the start of fragment B, we see that B starts with "AGCTTGCATG", so the primer "b" must include regions that can complement this segment.
Primer in Option 2 is designed to ensure that it binds effectively to the end of fragment A and prepares for the continuation into fragment B. The specificity and complementary nature of the primer to its target are crucial for successful amplification and subsequent processes, such as cloning or sequencing.