Correct option is A
Explanation-
Experimental Setup Summary:
The three conditions being compared for protein 'X':
1. Black: pH 7.0 → normal condition
2. Green: pH 3.0 → acidic
3. Red: pH 7.0 + 6 M guanidine hydrochloride → denaturing condition
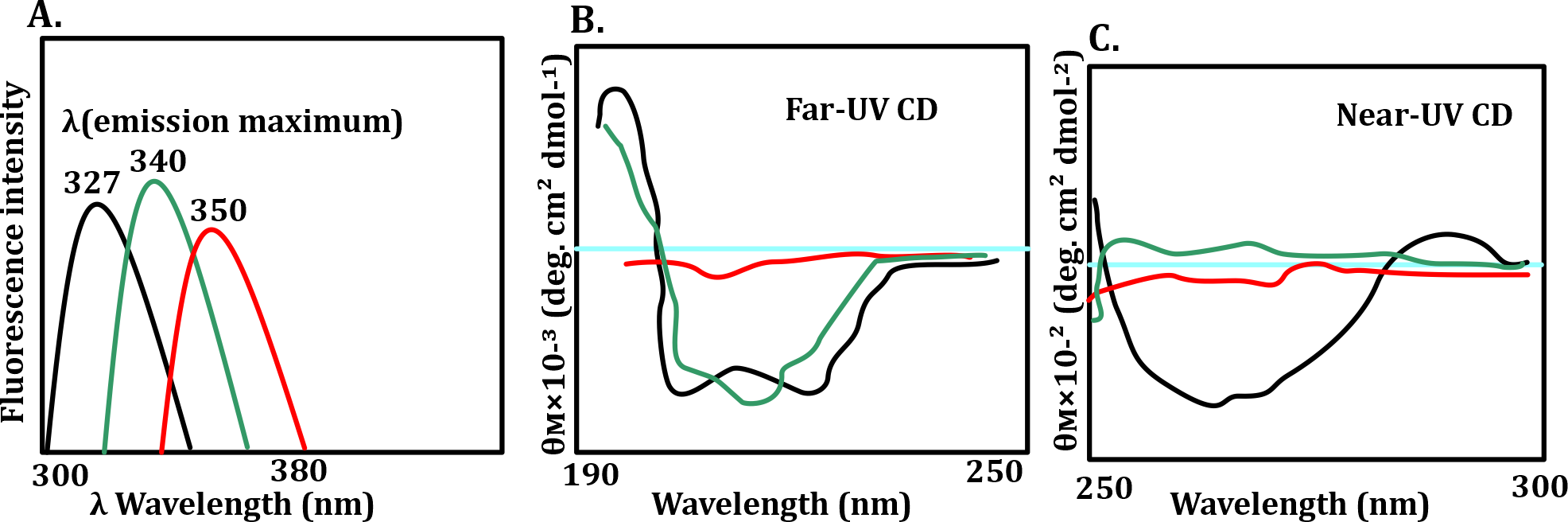
Panel A: Intrinsic Fluorescence (Tertiary Structure)
Fluorescence emission max for tryptophan:
327 nm (black) = buried in hydrophobic core → compact/folded
340 nm (green) = partial exposure → partially unfolded (molten globule)
350 nm (red) = fully exposed → unfolded
Interpretation: At pH 7.0, protein is folded.
At pH 3.0, partial unfolding → molten globule state.
With guanidine, fully unfolded.
Panel B: Far-UV CD (Secondary Structure)
Strong signals at pH 7.0 (black) → high secondary structure content.
Moderate signal at pH 3.0 (green) → partial structure.
Very low/flat signal with guanidine (red) → loss of secondary structure.
Interpretation:
Secondary structure intact at pH 7.0 , reduced at pH 3.0, destroyed with guanidine.
Panel C: Near-UV CD (Tertiary Structure)
Black (pH 7.0) → clear signal = well-packed tertiary structure.
Green (pH 3.0) → flat = molten globule, tertiary structure lost.
Red (guanidine) → flat = fully unfolded.
Interpretation:
Tertiary structure intact at pH 7.0 , disrupted at pH 3.0 and with guanidine.
Correct Answer: Option a - “Protein is fully folded at pH 7.0, acid-induced molten globule at pH 3.0 and unfolded in 6M guanidine hydrochloride.
Fluorescence: 327 → 340 → 350 nm (increasing exposure)
Far-UV CD: high → reduced → minimal (secondary structure loss)
Near-UV CD: intact → lost → lost (tertiary structure loss)
Incorrect options-
Option b: Says secondary structure is reduced at pH 7.0 — false, it's highest at pH 7.0.
Option c: Talks about hydrodynamic radius — not measured in this experiment.
Option d: Says extensive denaturation at both pH 3.0 and guanidine — wrong, pH 3.0 causes partial unfolding, not full.