Correct option is A
1. Denaturation of Proteins:
- Urea: A chaotropic agent that disrupts hydrogen bonds, leading to protein unfolding. It is often used to denature proteins by weakening the interactions between the polypeptide chains.
- Sodium dodecyl sulfate (SDS): Anionic detergent that binds to proteins and disrupts hydrophobic interactions, leading to the denaturation and unfolding of proteins.
- Guandinium hydrochloride (GnHCl): A strong chaotropic agent that is commonly used to denature proteins by disrupting both hydrogen bonds and ionic interactions.
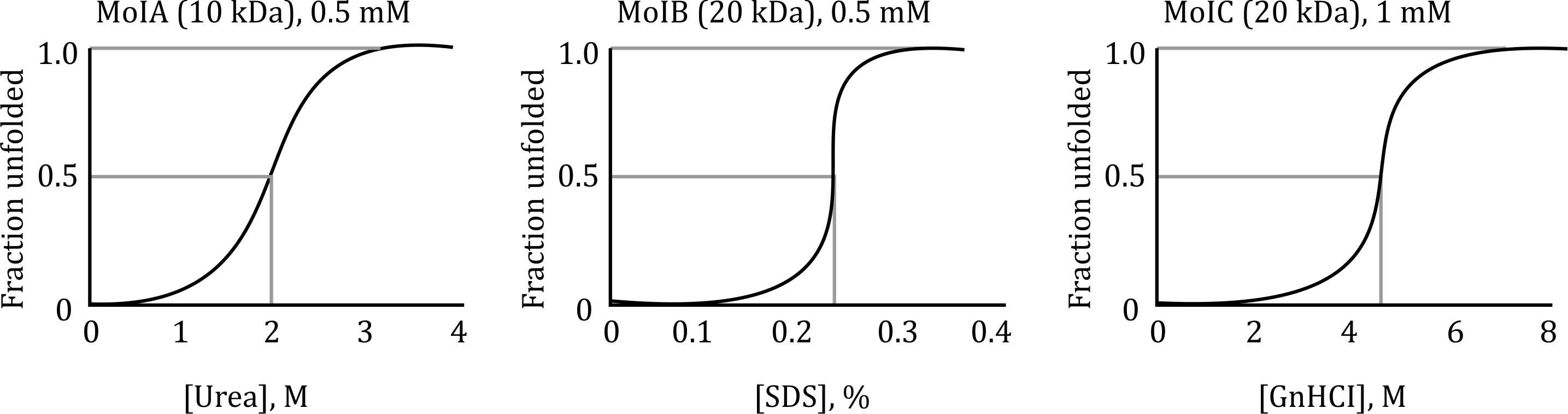
These denaturants unfold proteins in a concentration-dependent manner. The fraction of unfolded protein increases as the concentration of these agents increases.
2. Molecular Size and Denaturation:
- Larger proteins like MolC (20 kDa) require higher concentrations of denaturants to unfold compared to smaller proteins like MolA (10 kDa).
- The protein-to-RNA ratio in a denaturation study gives insight into the stability of the protein under the given denaturant concentrations.
3. Equilibrium of Folded and Unfolded Proteins:
- The equilibrium between the folded and unfolded states of proteins is influenced by the denaturant concentration.
- The condition where the number of folded molecules of protein is equal means that the fraction of unfolded protein is at a point where it’s balanced across different proteins, i.e., the unfolded fraction is similar for MolA, MolB, and MolC.
Option 1 Is Correct:
0.2 M urea; 0.05 % SDS; 4.5 M GnHCl:
- 0.2 M urea: This is an optimal concentration to denature MolA (10 kDa) and MolB (20 kDa) without overly affecting the folding balance. Urea concentrations higher than 0.2 M would overly destabilize the proteins, especially MolA.
- 0.05% SDS: This concentration is sufficient to denature proteins without excessively unfolding them, ensuring that MolA and MolB are in a comparable denaturation state to MolC.
- 4.5 M GnHCl: GnHCl is needed in higher concentrations for MolC (the larger protein), as it has a higher threshold for unfolding. This concentration ensures that MolC is properly denatured but does not cause excessive denaturation for the smaller proteins like MolA and MolB.
This combination ensures that all three proteins are in a comparable denaturation state, and the number of folded molecules of each protein will be equal.
Incorrect Options:
Option 2 (2 M urea; 0.05 % SDS; 1 M GnHCl):
- 2 M urea is too high and will over-denature MolA and MolB, leading to unequal folding between the proteins.
- 1 M GnHCl is insufficient to denature MolC effectively, leading to an imbalance in the degree of unfolding between proteins.
Option 3 (0.2 M urea; 0.25 % SDS; 4.5 M GnHCl):
- 0.25% SDS is too high, which would over-denature MolA and MolB and disrupt the balance between the proteins' folding states.
- While the urea and GnHCl concentrations are reasonable, the SDS concentration would cause excessive unfolding, making this combination unsuitable.
Option 4 (2 M urea; 0.25 % SDS; 1 M GnHCl):
- 2 M urea and 0.25% SDS would cause over-denaturation of MolA and MolB, leading to an imbalance in the protein folding.
- 1 M GnHCl is not enough to unfold MolC sufficiently, resulting in uneven denaturation between the three proteins.