Correct option is D
Explanation:
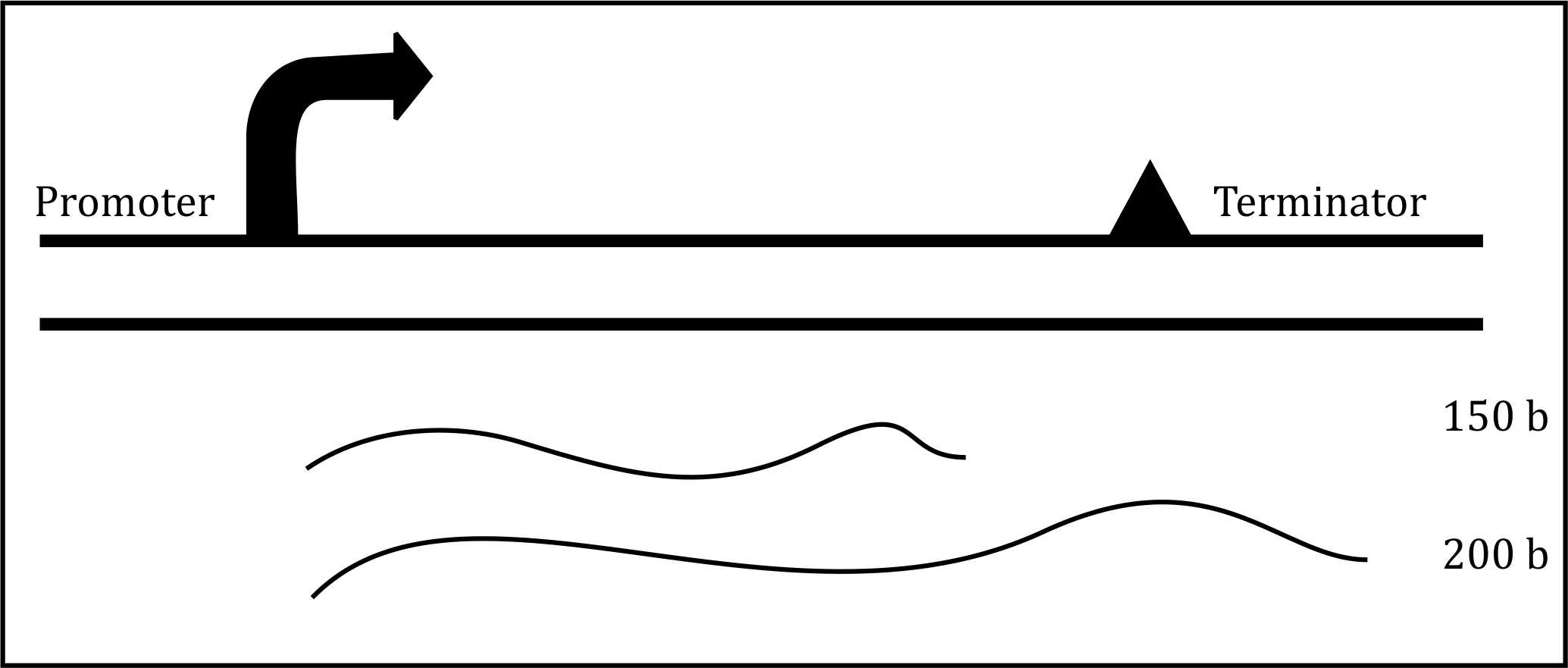
Understanding Rho-independent Termination Mechanism:
- Rho-independent termination relies on two key features:
- GC-rich stem-loop structure (hairpin) in the mRNA.
- A string of uracils (U) following the hairpin.
- Mechanism:
- The hairpin structure causes RNA polymerase to pause.
- The weak U-A interactions (between the transcript and template DNA) allow RNA to dissociate, ending transcription.
- Disrupting either the hairpin or the U-rich sequence can prevent termination, leading to a longer transcript.
Analysis of Each Mutation:
(A) Three nucleotides of the string of 8Ts were replaced by GCC.
- Effect on termination:
- The string of uracils is crucial for proper termination because U-A bonds are weak.
- Replacing three uracils with G/C bases strengthens base pairing, making termination inefficient.
- Expected transcript size:
- Long transcript (200 bases) because termination is disrupted.
(B) The 8T sequence was transferred to the template strand.
- Effect on termination:
- Normally, Ts in the non-template strand result in U’s in the mRNA transcript.
- If Ts are moved to the template strand, the transcript will have A’s instead of U’s.
- A-rich sequences do not cause termination, as A-U pairs are stronger than U-A pairs.
- Expected transcript size:
- Long transcript (200 bases) because the U-run is missing, preventing termination.
(C) Disrupting the hairpin structure.
- Effect on termination:
- The hairpin structure is necessary to pause RNA polymerase.
- If hairpin formation is disrupted, termination efficiency drops significantly.
- Expected transcript size:
- Long transcript (200 bases) because RNA polymerase fails to pause.
(D) Disrupting the hairpin but restoring it with compensatory mutations.
- Effect on termination:
- Compensatory mutations restore base pairing in the hairpin.
- Since the hairpin functions properly, termination will occur as expected.
- Expected transcript size:
- Short transcript (150 bases) because termination is restored.
Information Booster:
- Rho-independent termination depends on the formation of a GC-rich hairpin followed by U-rich sequences.
- U-A interactions are weaker than G-C interactions, making the termination site susceptible to disruption.
- Mutation in the U-run region can lead to transcription readthrough, increasing transcript length.
- Disrupting the hairpin structure prevents RNA polymerase pausing, causing failure in termination.
- Compensatory mutations restoring the hairpin structure can recover proper termination efficiency.
- Mutations in bacterial terminators can lead to antibiotic resistance by altering gene expression levels.