Correct option is A
The question revolves around understanding the recognition sequences and cleavage sites of the given restriction enzymes, and matching vector fragments with insert fragments that create compatible cohesive ends for ligation.
EcoRI recognizes G^AATTC, cutting between G and A, producing sticky ends.
HincII recognizes GTY^RAC, cutting within the sequence, producing blunt ends.
EcoRV recognizes GAT^ATC, producing blunt ends.
BamHI recognizes G^GATCC, producing sticky ends.
BglII recognizes A^GATCT, producing sticky ends compatible with BamHI ends.
The key to this problem is that fragments produced by enzymes that create compatible sticky ends can be ligated successfully.
EcoRI sticky ends are compatible with EcoRI sticky ends only.
BamHI and BglII generate compatible sticky ends (both generate 5' overhangs of GATC).
HincII and EcoRV produce blunt ends, which can ligate with each other but less efficiently.
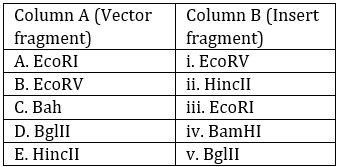
Looking at the options and the table:
Option B pairs EcoRV (vector fragment) with HincII (insert fragment) - both blunt ends, compatible.
D pairs BglII with BamHI - compatible sticky ends.
E pairs HincII with BglII is incorrect because HincII produces blunt ends, and BglII sticky ends are not compatible with blunt ends.
A pairs EcoRI with EcoRV (sticky with blunt) - not compatible.
C pairs BamHI with EcoRI (sticky ends with different overhangs) - incompatible.
Therefore, option (b) is the only one with correct compatible ends.