Correct option is D
To determine the primary structure of the peptide, we analyze the given experimental data.
Step 1: Acid Hydrolysis (Complete Breakdown)
- The given set of amino acids: Ala, Arg, Cys, Glx, Gly, Lys, Leu, Met, Phe, Thr.
- This confirms all amino acids present but does not give sequence information.
Step 2: Aminopeptidase M
- No fragments were obtained, indicating that the N-terminus is blocked, possibly due to cyclization.
Step 3: Carboxypeptidase A + B
- No fragments, meaning that the C-terminus is blocked, further supporting cyclization.
Step 4: Trypsin Digestion + Edman Degradation
- Trypsin cleaves at Lys (K) and Arg (R).
- Two fragments obtained:
- Cys-Gly-Leu-Phe-Arg (Fragment 1)
- Thr-Ala-Met-Gln-Lys (Fragment 2)
From these results:
- Since Trypsin cleaves after Arg and Lys, we deduce that Arg is at the C-terminal of one fragment, and Lys is at the C-terminal of the other.
- However, since Aminopeptidase and Carboxypeptidase gave no results, the peptide is most likely cyclic.
Final Answer:
The cyclic nature of the peptide is supported by:
- No free N-terminal or C-terminal (Step 2 and Step 3 results).
- Matching sequence fragments with the cyclic representation.
Thus, the correct answer is:(4)
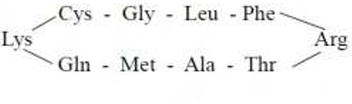
Explanation:
- A cyclic peptide means that the amino acids form a closed-loop structure rather than a linear sequence.
- Since both trypsin digestion products match the sequence in the given cyclic structure, the primary structure must be cyclic.
- Linear options (1, 2, 3) are ruled out due to missing free ends.







